Day 333: Sneaky shape-shifting molecule mimics DNA to trick viruses

25th April 2015
Author: Geoff Maitland, IChemE President 2014–2015.
Very few discoveries truly revolutionise the way we look at the world.
However, the discovery of the structure of DNA is one of them. And it was on this day in 1953, that the structure of DNA was published in the journal Nature.
This work was published at the same time in a number of papers in Nature by three teams: Watson and Crick; Wilkins, Stokes, and Wilson; and Franklin and Gosling.
The key break through for Watson and Crick's work came from Rosalind Franklin who studied DNA using X-ray crystallography, but this was largely unacknowledged at the time. In 1962 Crick and Watson, along with Wilkins, received a Nobel Prize for their discovery. Rosalind had died four years earlier so was not eligible for a Nobel Prize.
So to ensure that we celebrate all their work today, I thought I would bring to your attention a recent innovation, which would not have been possible without this major discovery.
A team of scientists and engineers from the University of Chicago (UChicago) and the Massachusetts Institute of Technology (MIT), US, have developed a new spectroscopy method that could prove useful in developing the next generation of anti-viral treatments.
The team used synthetically designed shape-shifting molecules which are able to resemble natural DNA bases, but can convert into a different molecular structure by repositioning their hydrogen atoms on nitrogen and oxygen atoms.

These shape-shifting structures are called tautomers, and the idea for them also came from Watson and Crick in 1953, who announced this idea with their description of the structure of DNA.
This spectroscopy method improves our understanding of the molecular process by which an anti-HIV drug (KP1212) causes lethal mutations in the virus’ genetic material.
Viruses are so successful due to their ability to rapidly mutate to environmental pressures. This is what makes them so resistant to anti-viral drugs. But research has shown that it is possible to kill viruses using lethal mutagenesis.
The lethal mutagenesis process works by forcing viruses to mutate at a rate so high, they are unable to properly manage their genetic material.
Andrei Tokmakoff, the Henry G. Gale Distinguished Service Professor in Chemistry at UChicago, said: “In order to make this work, you need a stealth mutagen. You need something sneaky, something that the virus isn’t going to recognise as a problem.”
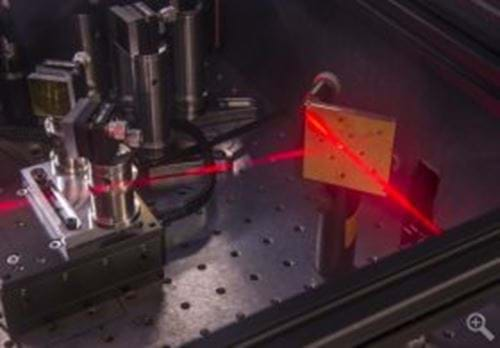
This data was collected with two-dimensional infrared spectroscopy, an advanced laser technique that combines ultra fast time resolution with high sensitivity to chemical structure.
John Essigmann, MIT’s William and Betsy Leitch Professor of Chemistry, Toxicology and Biological Engineering, explained: “Two-dimensional infrared spectroscopy will be critical on the path ahead. It lets us look at the structures that exist in aqueous solution, which is the natural milieu of cells."
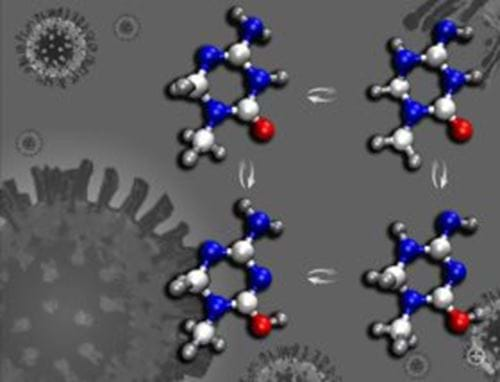
The team used lethally mutagenic molecules that resemble natural DNA bases, the adenine-thymine, cytosine-guanine base pairs.
Sam Peng, the lead author in this study and a visiting graduate research assistant at UChicago, said: “These analogs can bind to the wrong base partners and therefore lead to genetic mutations.”
For example, while in nature, cytosine will only form a pair bond with guanine, the team were able to use anti-HIV drug KP1212 to induce a mutation to cause cytosine to pair with adenine.
It is suggested that KP1212 derives its mutagenicity by shape shifting - it is a tautomer. “The shuffled hydrogen positions in rare tautomers alter the hydrogen bonding patterns, resulting in incorrect base paring,” said Sam.
Sam, Andrei, John and their colleagues reported their findings in the Proceedings of the National Academy of Sciences under the title: ‘Two-dimensional IR spectroscopy of the anti-HIV agent KP1212 reveals protonated and neutral tautomers that influence pH-dependent mutagenicity’.
Most experimental tools would have difficulty distinguishing between the normal and shape-shifted molecules, however, two-dimensional infrared spectroscopy enabled the researchers to distinguish between the structures. The team were also able measure how fast the shape-shifting occurred – in 20 billionths of a second.
This impressive work shows the importance of collaboration between different fields, and demonstrates the power of the discovery of DNA. It also has important applications for future anti-viral drugs, especially those designed to fight HIV.
If you are chemical engineer working with DNA why not get in touch and tell us how you were inspired?
ChemEng365 blog
Geoff Maitland launched this blog during his IChemE presidency in 2014. ChemEng365 features 365 chemical engineering successes and achievements throughout his year-long presidency.